Workshop
NEON Brownbag: Intro to HDF5 at NEON
NEON
This NEON internal brownbag introduces the concept of Hierarchical Data Formats in the context of developing the NEON HDF5 operational file format. Look here to discover resources on HDF5, code snippets in R, Python and Matlab to use H5 files and some example H5 files for Remote Sensing Hyperspectral data and time series temperature data.
Things to do before the workshop
Please review, download and setup the following, prior to attending the brownbag.
###Data to Download
[[nid:6329]] [[nid:6330]]
Download the Free H5 Viewer
The free H5 viewer will allow you to explore H5 data, using a graphic interface. HDF5 viewer can be downloaded from this page. More details on how to set up HDF5Viewer are at the end of this page.
Background Information
Schedule
| Time | Topic |
|---|---|
| 12:00 | Hand-on exploration of the HDF5 Data Format |
| 12:20 | Working with HDF5 in Python - live demo. |
| 12:30 | NEON HDF5 Format - what's next |
Instructors
- David Hulslander
- Josh Elliot
- Leah A. Wasser
- Tristan Goulden
Additional Set Up Instructions
Install HDFView
The free HDFView application allows you to explore the contents of an HDF5 file.
To install HDFView:
-
Click to go to the download page.
-
From the section titled HDF-Java 2.1x Pre-Built Binary Distributions select the HDFView download option that matches the operating system and computer setup (32 bit vs 64 bit) that you have. The download will start automatically.
-
Open the downloaded file.
- Mac - You may want to add the HDFView application to your Applications directory.
- Windows - Unzip the file, open the folder, run the .exe file, and follow directions to complete installation.
- Open HDFView to ensure that the program installed correctly.
| Time | Topic |
|---|---|
| 12:00 | Hand-on exploration of the HDF5 Data Format |
| 12:20 | Working with HDF5 in Python - live demo. |
| 12:30 | NEON HDF5 Format - what's next |
Hierarchical Data Formats - What is HDF5?
Learning Objectives
After completing this tutorial, you will be able to:
- Explain what the Hierarchical Data Format (HDF5) is.
- Describe the key benefits of the HDF5 format, particularly related to big data.
- Describe both the types of data that can be stored in HDF5 and how it can be stored/structured.
About Hierarchical Data Formats - HDF5
The Hierarchical Data Format version 5 (HDF5), is an open source file format that supports large, complex, heterogeneous data. HDF5 uses a "file directory" like structure that allows you to organize data within the file in many different structured ways, as you might do with files on your computer. The HDF5 format also allows for embedding of metadata making it self-describing.

Hierarchical Structure - A file directory within a file
The HDF5 format can be thought of as a file system contained and described
within one single file. Think about the files and folders stored on your computer.
You might have a data directory with some temperature data for multiple field
sites. These temperature data are collected every minute and summarized on an
hourly, daily and weekly basis. Within one HDF5 file, you can store a similar
set of data organized in the same way that you might organize files and folders
on your computer. However in a HDF5 file, what we call "directories" or "folders"
on our computers, are called groups and what we call files on our
computer are called datasets.
2 Important HDF5 Terms
- Group: A folder like element within an HDF5 file that might contain other groups OR datasets within it.
- Dataset: The actual data contained within the HDF5 file. Datasets are often (but don't have to be) stored within groups in the file.
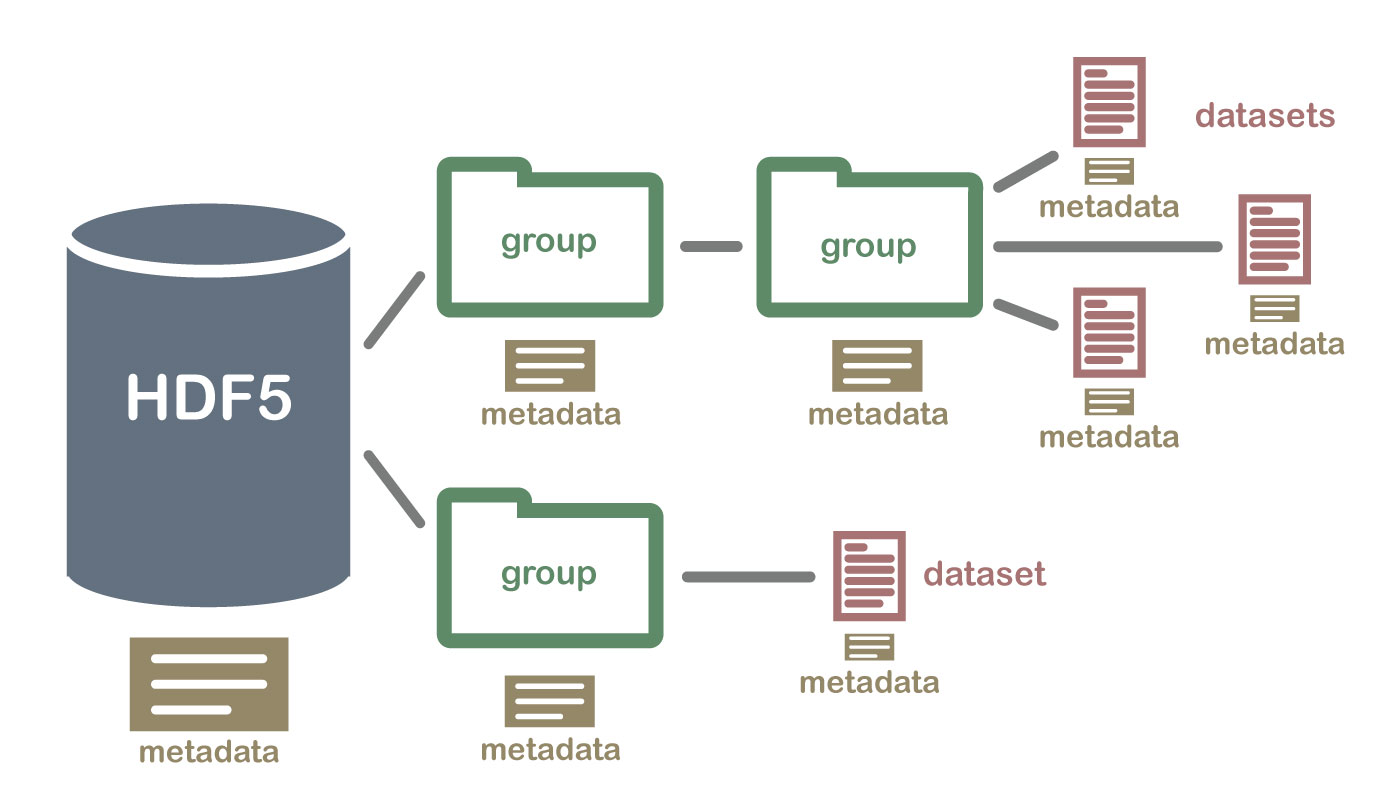
An HDF5 file containing datasets, might be structured like this:
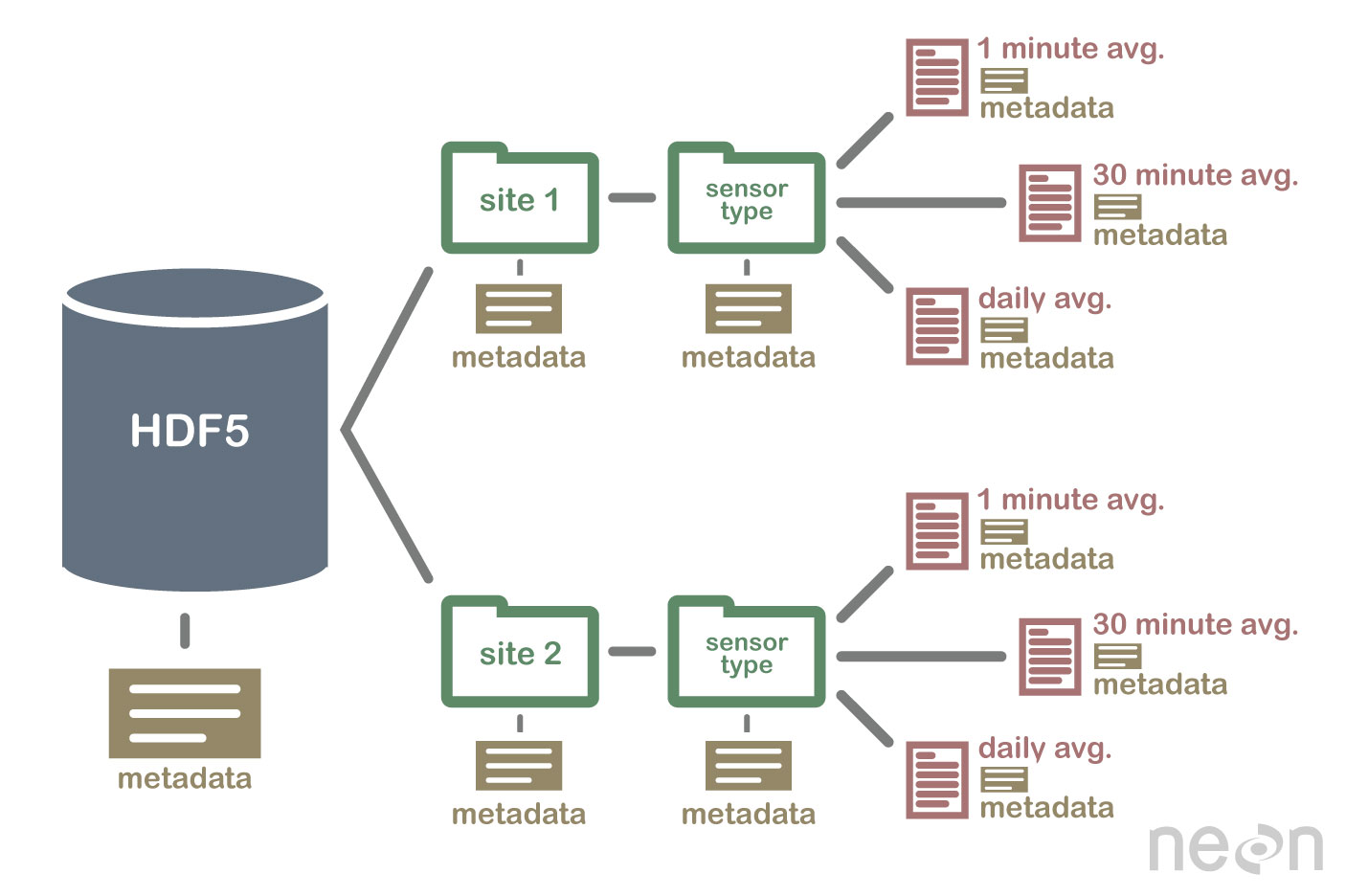
HDF5 is a Self Describing Format
HDF5 format is self describing. This means that each file, group and dataset can have associated metadata that describes exactly what the data are. Following the example above, we can embed information about each site to the file, such as:
- The full name and X,Y location of the site
- Description of the site.
- Any documentation of interest.
Similarly, we might add information about how the data in the dataset were collected, such as descriptions of the sensor used to collect the temperature data. We can also attach information, to each dataset within the site group, about how the averaging was performed and over what time period data are available.
One key benefit of having metadata that are attached to each file, group and
dataset, is that this facilitates automation without the need for a separate
(and additional) metadata document. Using a programming language, like R or
Python, we can grab information from the metadata that are already associated
with the dataset, and which we might need to process the dataset.
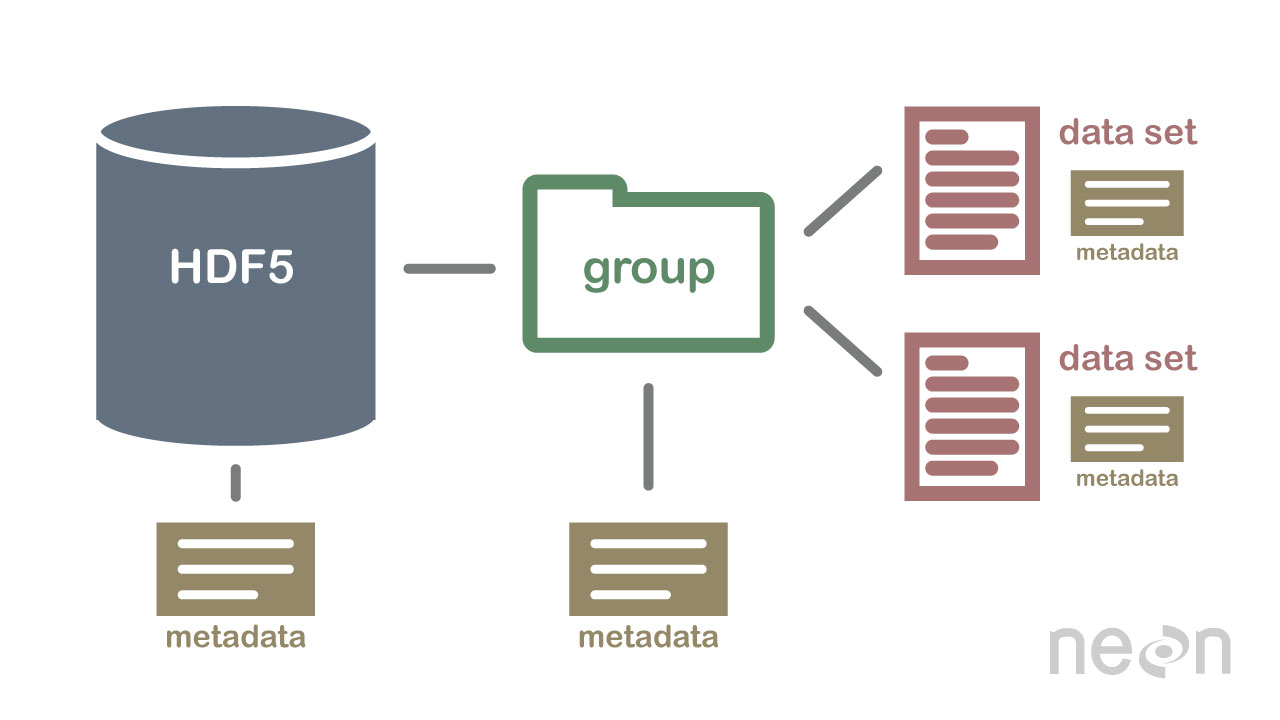
Compressed & Efficient subsetting
The HDF5 format is a compressed format. The size of all data contained within
HDF5 is optimized which makes the overall file size smaller. Even when
compressed, however, HDF5 files often contain big data and can thus still be
quite large. A powerful attribute of HDF5 is data slicing, by which a
particular subsets of a dataset can be extracted for processing. This means that
the entire dataset doesn't have to be read into memory (RAM); very helpful in
allowing us to more efficiently work with very large (gigabytes or more) datasets!
Heterogeneous Data Storage
HDF5 files can store many different types of data within in the same file. For example, one group may contain a set of datasets to contain integer (numeric) and text (string) data. Or, one dataset can contain heterogeneous data types (e.g., both text and numeric data in one dataset). This means that HDF5 can store any of the following (and more) in one file:
- Temperature, precipitation and PAR (photosynthetic active radiation) data for a site or for many sites
- A set of images that cover one or more areas (each image can have specific spatial information associated with it - all in the same file)
- A multi or hyperspectral spatial dataset that contains hundreds of bands.
- Field data for several sites characterizing insects, mammals, vegetation and meteorology.
- A set of images that cover one or more areas (each image can have unique spatial information associated with it)
- And much more!
Open Format
The HDF5 format is open and free to use. The supporting libraries (and a free
viewer), can be downloaded from the
HDF Group
website. As such, HDF5 is widely supported in a host of programs, including
open source programming languages like R and Python, and commercial
programming tools like Matlab and IDL. Spatial data that are stored in HDF5
format can be used in GIS and imaging programs including QGIS, ArcGIS, and
ENVI.
Summary Points - Benefits of HDF5
- Self-Describing The datasets with an HDF5 file are self describing. This allows us to efficiently extract metadata without needing an additional metadata document.
- Supporta Heterogeneous Data: Different types of datasets can be contained within one HDF5 file.
- Supports Large, Complex Data: HDF5 is a compressed format that is designed to support large, heterogeneous, and complex datasets.
- Supports Data Slicing: "Data slicing", or extracting portions of the dataset as needed for analysis, means large files don't need to be completely read into the computers memory or RAM.
-
Open Format - wide support in the many tools: Because the HDF5 format is
open, it is supported by a host of programming languages and tools, including
open source languages like R and
Pythonand open GIS tools likeQGIS.
HDFView: Exploring HDF5 Files in the Free HDFview Tool
In this tutorial you will use the free HDFView tool to explore HDF5 files and the groups and datasets contained within. You will also see how HDF5 files can be structured and explore metadata using both spatial and temporal data stored in HDF5!
Learning Objectives
After completing this activity, you will be able to:
- Explain how data can be structured and stored in HDF5 format.
- Navigate to metadata in an HDF5 file, making it "self describing".
- Explore HDF5 files using the free HDFView application.
Tools You Will Need
Install the free HDFView application. This application allows you to explore the contents of an HDF5 file easily. Click here to go to the download page.
Data to Download
Download NEON Imaging Spectrometer Data at SJER (2024) - NEON_D17_SJER_DP3_254000_4108000_bidirectional_reflectance.h5
These hyperspectral remote sensing data provide information on the National Ecological Observatory Network's San Joaquin Exerimental Range field site. The data were collected over the San Joaquin field site located in California (Domain 17) and processed at NEON headquarters. The entire dataset can be accessed from the Spectrometer orthorectified surface bidirectional reflectance - mosaic page on the NEON data portal.
Download NEON Eddy Covariance Data at SJER (2024-04-01)
The SAE data were collected by the National Ecological Observatory Network's flux towers at field sites across the US. The entire dataset can be accessed from the Bundled data products - eddy covariance page on the NEON data portal.
Download Eddy Covariance DatasetInstalling HDFView
Select the HDFView download option that matches the operating system (Mac OS X, Windows, or Linux) and computer setup (32 bit vs 64 bit) that you have.
Hierarchical Data Format 5 - HDF5
Hierarchical Data Format version 5 (HDF5), is an open file format that supports large, complex, heterogeneous data. Some key points about HDF5:
- HDF5 uses a "file directory" like structure.
- The HDF5 data models organizes information using
Groups. Each group may contain one or moredatasets. - HDF5 is a self describing file format. This means that the metadata for the data contained within the HDF5 file, are built into the file itself.
- One HDF5 file may contain several heterogeneous data types (e.g. images, numeric data, data stored as strings).
For more introduction to the HDF5 format, see our About Hierarchical Data Formats - What is HDF5? tutorial.
In this tutorial, we will explore two different types of data saved in HDF5. This will allow us to better understand how one file can store multiple different types of data, in different ways.
Part 1: Exploring Hyperspectral Imagery stored in HDF5

First, we will explore a hyperspectral dataset, collected by the NEON Airborne Observation Platform (AOP) and saved in HDF5 format. In the hyperpsectral data cubes, each pixel in the dataset contains reflectance values for hundreds of bands (426) collected by the sensor.
A few notes about hyperspectral imagery:
- An imaging spectrometer, which collects hyperspectral imagery, records light energy reflected off objects on the earth's surface.
- The data are inherently spatial. Each pixel in the image is located spatially and represents an area of ground on the earth.
- Similar to an RGB (Red, Green, Blue) camera, an imaging spectrometer records reflected light energy. Each pixel contain several hundred bands of reflectance data.
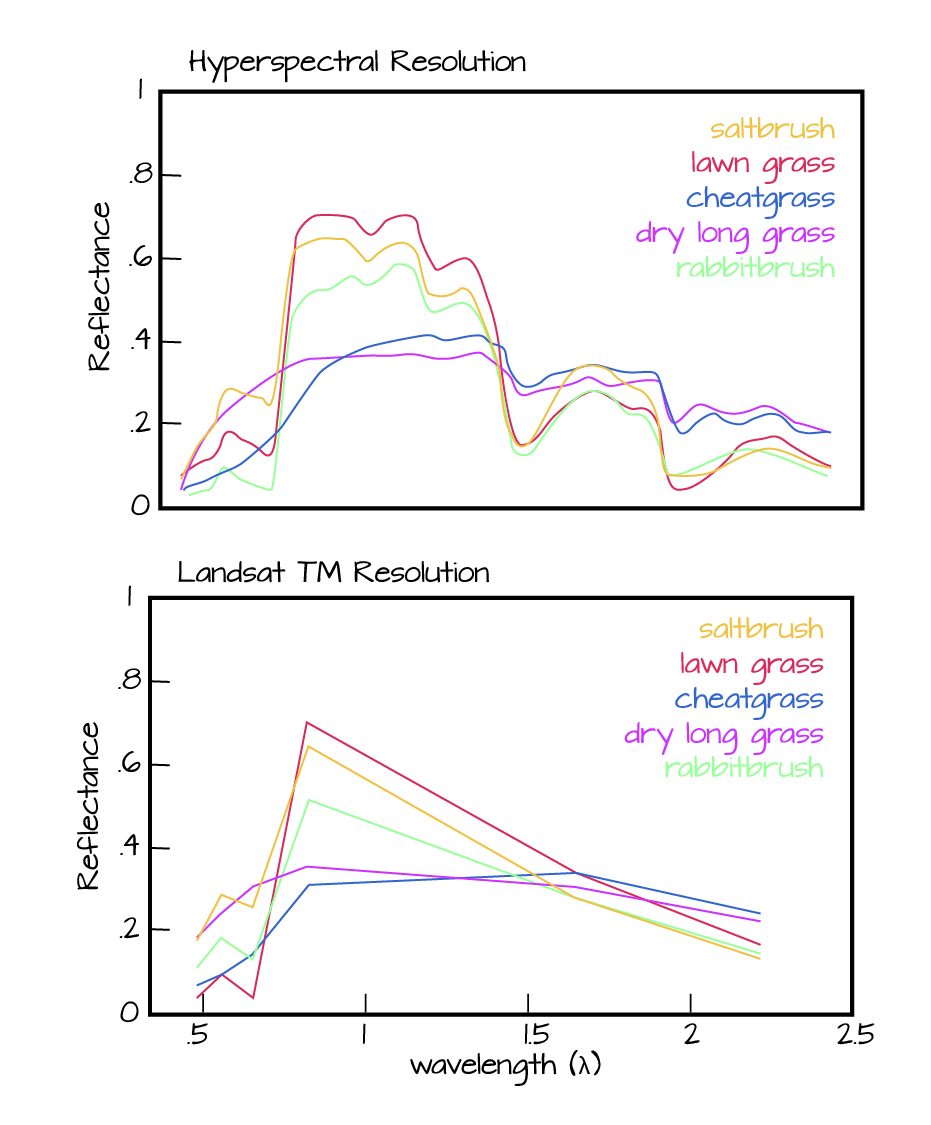
Read more about hyperspectral remote sensing data:
- About Hyperspectral Remote Sensing Data tutorial on this site.
Let's open some hyperspectral imagery stored in HDF5 format to see what the file structure can like for a different type of data.
Open the Reflectance H5 file in HDFView
To begin, open the HDFView application.
Within the HDFView application, select File --> Open and navigate to the folder
where you saved the NEON_D17_SJER_DP3_254000_4108000_bidirectional_reflectance.h5 file on your computer. Open this file in HDFView.
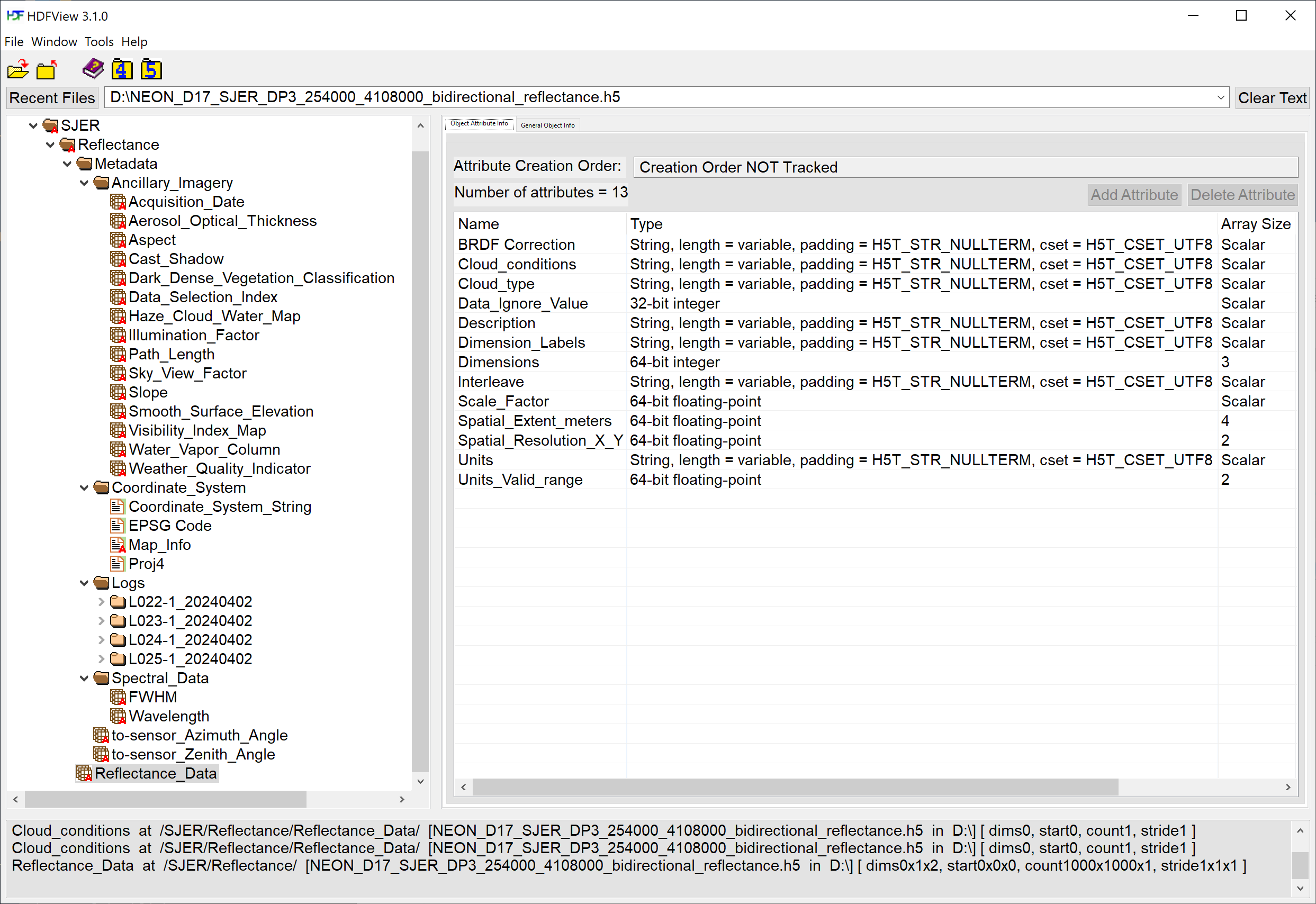
Open the file and expand the sub-folders. This file is composed of a Reflectance dataset (called Reflectance_Data) along with additional Metadata containing the following sub-folders:
-
Ancillary_Imagery: Datasets including ATCOR inputs and other Quality indicators such as theWeather_Quality_Indicator, containing information about the cloud conditions during the flight (for each pixel). -
Coordinate_System: geographic information for the dataset. -
Logs: Log files for each flight line containing ATCOR processing information and inputs, BRDF correction parameters, and the solar azimuth and zenith angles. -
Spectral_Data: Full Width Half Max (FWHM) and Wavelength for each of the 426 spectral bands.
Let's first look at the metadata stored in the Coordinate_System folder. This group contains all of the spatial information that a GIS program would need to project the data spatially.
Next, double click on the Wavelength dataset. Note that this dataset contains the central wavelength value for each band in the dataset.
Finally, click on the Reflectance_Data dataset. Note that in the metadata for the dataset that the structure of the dataset is 426 x 1000 x 1000 (wavelength, x, y), as indicated in the metadata. Right click on the reflectance dataset and select Open As. Click Image in the "display as" settings on the left hand side of the popup.
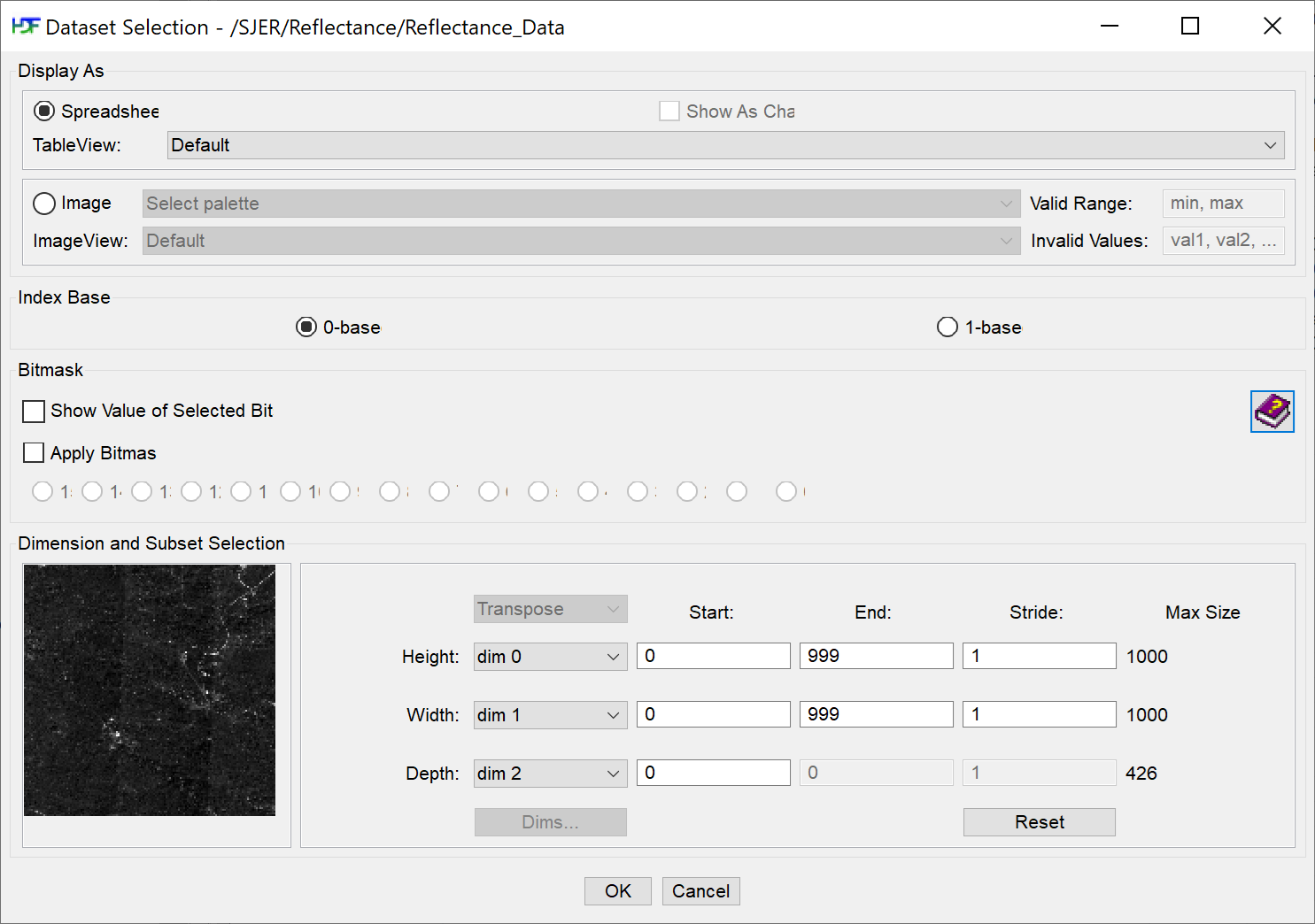
Notice an image preview appears on the left of the pop-up window. Click OK to open the image. You may have to play with the brightness and contrast settings in the viewer to see the data properly.

Explore the spectral dataset in the HDFViewer taking note of the metadata and data stored within the file.
Part 2: Exploring Surface Atmosphere Exchange (SAE) Data in HDFView
Next, we will look at the SAE bundled eddy covariance h5 data. As in the first part, we will start by opening the h5 file (download from the link at the top of this tutorial) in the viewer to get a better idea of how this data is structured.
Open the Bundled Eddy Covariance H5 file in HDFView
Open the HDFView application. Within the application, select File --> Open and navigate to the folder where you saved the SAE hdf5 file on your computer. Open this file in HDFView.
If you click on the name of the HDF5 file in the left hand window of HDFView, you can view metadata for the file. This will be located in the bottom window of the application.
Explore File Structure in HDFView
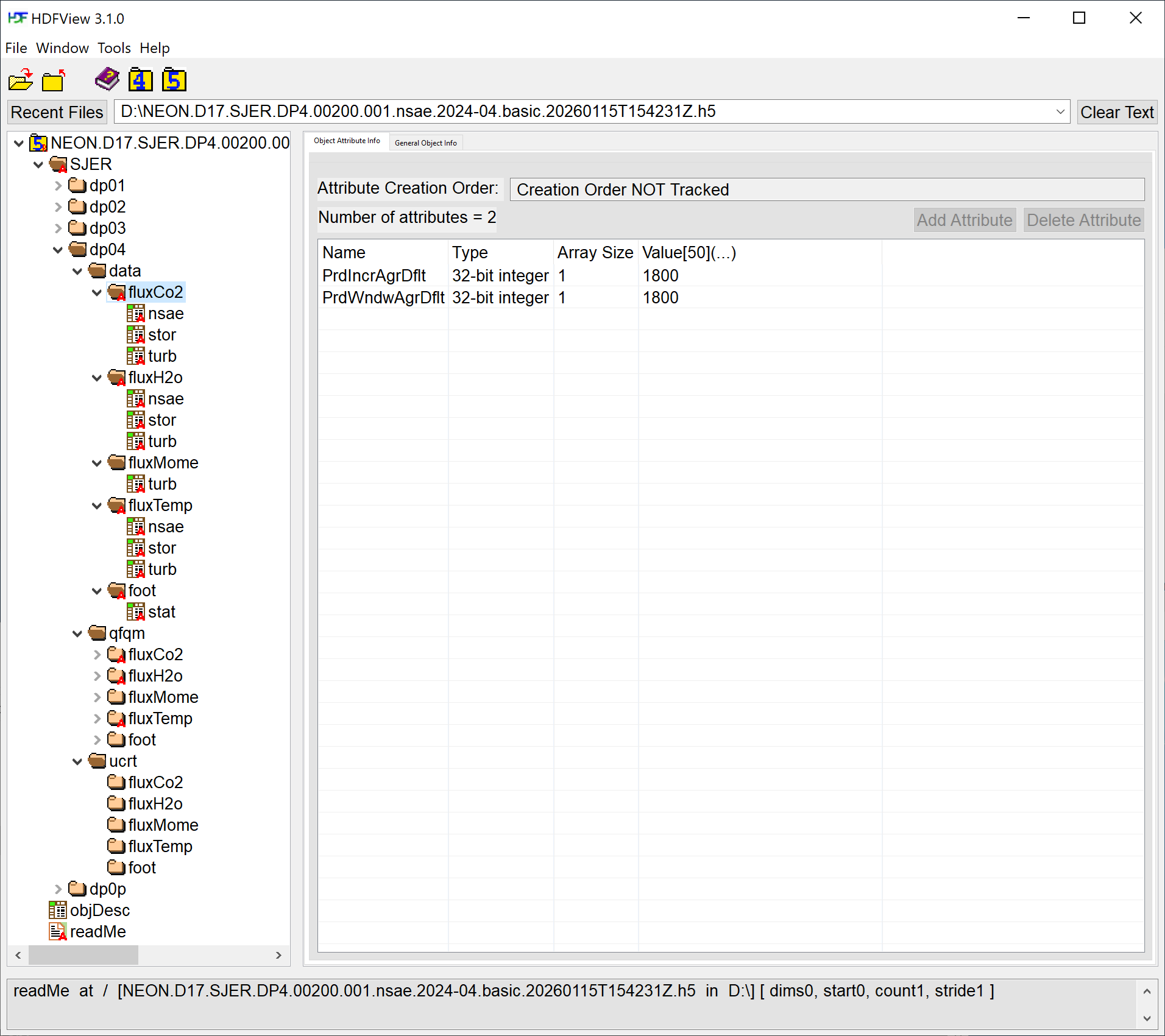
Next, explore the structure of this bundled eddy covariane file.
Notice at the bottom there is a readMe attribute. If you double click on this, you'll see the text "Net Surface Atmosphere Exchange (NSAE) HDF5 File Structure Description. The NSAE file you downloaded from NEON data portal is in the HDF5 format. This document describes the HDF5 file structure. This file will provide the HDF5 hierarchical layout of the file and a description of each HDF5 group level. The full descriptions of objects can be found in the objDesc data table provided within the HDF5 file. The 'Exploring NEON Eddy-Covariance Data Products in HDF5 file format' document provides a greater level of detail ..."
Documentation for each NEON data product is contained on the respective data product page. It is strongly recommended to peruse the relevant documentation, starting with the Quick Start Guides. The document referenced above in the readMe is linked here: Exploring NEON Eddy-Covariance Data Products in HDF5 file format .
Now that you've read the readMe, and referencing the document above, take a look at the structure of the data in HDFView.
Notice that there are multiple groups (folders) under the SJER root folder starting with dp. Expand these folders by double clicking on the folder icons. These represent the different data product levels, from 01 to 04, as well as level 0 prime.
-
dp01: Level 1 -
dp02: Level 2 -
dp03: Level 3 -
dp04: Level 4 -
dp0p: Level 0 prime
Under each of the levels there is a data folder with subfolders labeled by the data product identification codes as well as quality information (qfqm) and uncertainty (ucrt). Notice that there is also metadata associated with each group.
Within the dp04/data group there are five more groups: fluxCo2, fluxH2o, fluxMome, fluxTemp, and foot. What data are contained within these groups?

So this is another example of how a NEON HDF5 file is structured. Take some time to explore this HDF5 dataset within the HDFViewer, using the reference document as needed.