Tutorial
Raster 00: Intro to Raster Data in R
Authors: Leah A. Wasser, Megan A. Jones, Zack Brym, Kristina Riemer, Jason Williams, Jeff Hollister, Mike Smorul
Last Updated: Apr 2, 2026
In this tutorial, we will review the fundamental principles, packages and metadata/raster attributes that are needed to work with raster data in R. We discuss the three core metadata elements that we need to understand to work with rasters in R: CRS, extent and resolution. We also explore missing and bad data values as stored in a raster and how R handles these elements. Finally, we introduce the GeoTiff file format.
Learning Objectives
After completing this tutorial, you will be able to:
- Understand what a raster dataset is and its fundamental attributes.
- Know how to explore raster attributes in R.
- Be able to import rasters into R using the
terrapackage. - Be able to quickly plot a raster file in R.
- Understand the difference between single- and multi-band rasters.
Things You’ll Need To Complete This Tutorial
You will need the most current version of R and, preferably, RStudio loaded
on your computer to complete this tutorial.
Install R Packages
-
terra:
install.packages("terra") -
More on Packages in R – Adapted from Software Carpentry.
Download Data
Data required for this tutorial will be downloaded using neonUtilities in the lesson.
The LiDAR and imagery data used in this lesson were collected over the National Ecological Observatory Network's Harvard Forest (HARV) field site.
The entire dataset can be accessed from the NEON Data Portal.
Set Working Directory: This lesson will explain how to set the working directory. You may wish to set your working directory to some other location, depending on how you prefer to organize your data.
An overview of setting the working directory in R can be found here.
- Read more about the
terrapackage in R. - NEON Data Skills tutorial: Raster Data in R - The Basics
- NEON Data Skills tutorial: Image Raster Data in R - An Intro
About Raster Data
Raster or "gridded" data are stored as a grid of values which are rendered on a map as pixels. Each pixel value represents an area on the Earth's surface.
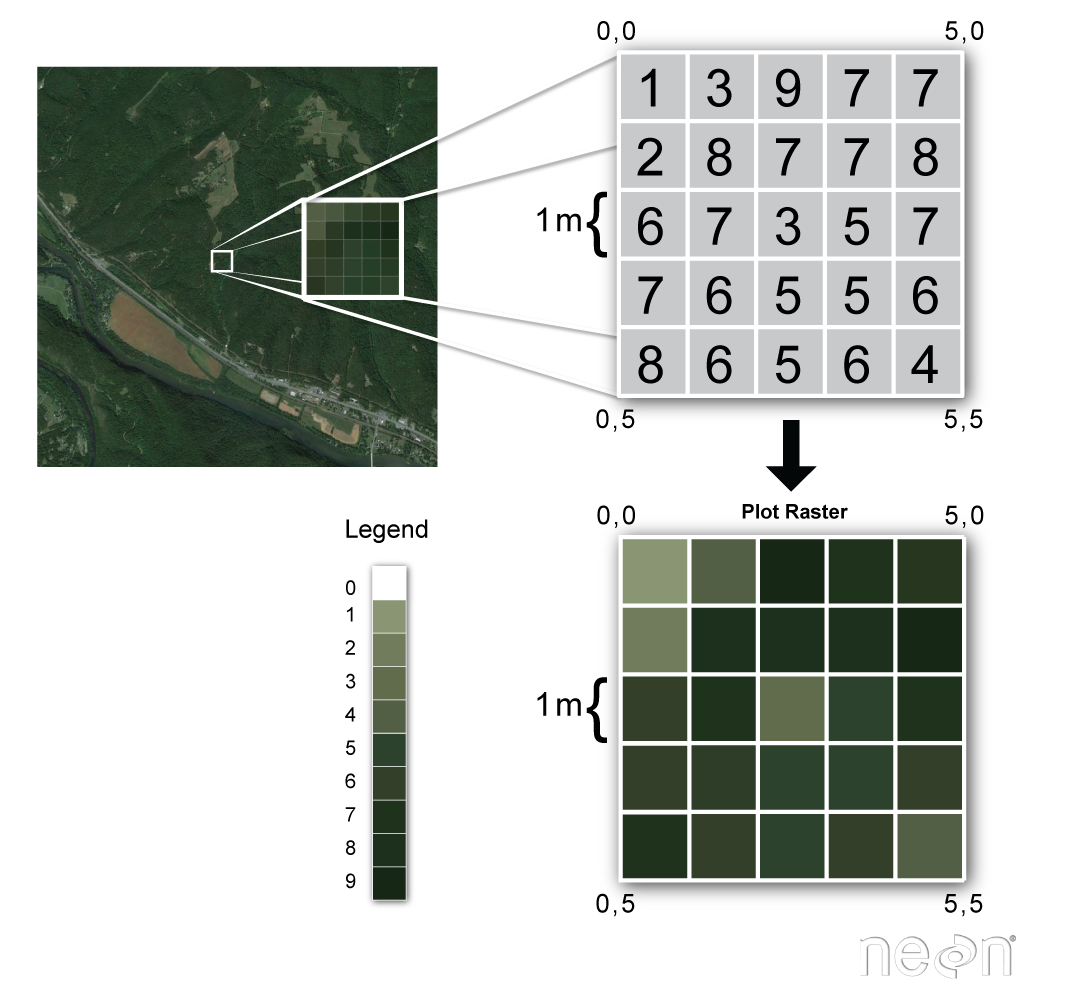

Raster Data in R
Let's first import a raster dataset into R and explore its metadata. To open rasters in R, we will use the terra package.
library(terra)
# set working directory, you can change this if desired
wd <- "~/data/"
setwd(wd)
Download LiDAR Raster Data
We can use the neonUtilities function byTileAOP to download a single elevation tiles (DSM and DTM). You can run help(byTileAOP) to see more details on what the various inputs are. For this exercise, we'll specify the UTM Easting and Northing to be (732000, 4713500), which will download the tile with the lower left corner (732000,4713000). By default, the function will check the size total size of the download and ask you whether you wish to proceed (y/n). This file is ~8 MB, so make sure you have enough space on your local drive. You can set check.size=TRUE if you want to check the file size before downloading.
byTileAOP(dpID='DP3.30024.001', # lidar elevation
site='HARV',
year='2022',
easting=732000,
northing=4713500,
check.size=FALSE, # set to TRUE or remove if you want to check the size before downloading
savepath = wd)
This file will be downloaded into a nested subdirectory under the ~/data folder, inside a folder named DP3.30024.001 (the Data Product ID). The file should show up in this location: ~/data/DP3.30024.001/neon-aop-products/2022/FullSite/D01/2022_HARV_7/L3/DiscreteLidar/DSMGtif/NEON_D01_HARV_DP3_732000_4713000_DSM.tif.
Open a Raster in R
We can use terra's rast("path-to-raster-here") function to open a raster in R.
Data Tip: VARIABLE NAMES! To improve code
readability, file and object names should be used that make it clear what is in
the file. The data for this tutorial were collected over from Harvard Forest (HARV),
sowe'll use the naming convention of DATATYPE_HARV.
# Load raster into R
dsm_harv_file <- paste0(wd, "DP3.30024.001/neon-aop-products/2022/FullSite/D01/2022_HARV_7/L3/DiscreteLidar/DSMGtif/NEON_D01_HARV_DP3_732000_4713000_DSM.tif")
DSM_HARV <- rast(dsm_harv_file)
# View raster structure
DSM_HARV
## class : SpatRaster
## dimensions : 1000, 1000, 1 (nrow, ncol, nlyr)
## resolution : 1, 1 (x, y)
## extent : 732000, 733000, 4713000, 4714000 (xmin, xmax, ymin, ymax)
## coord. ref. : WGS 84 / UTM zone 18N (EPSG:32618)
## source : NEON_D01_HARV_DP3_732000_4713000_DSM.tif
## name : NEON_D01_HARV_DP3_732000_4713000_DSM
## min value : 317.91
## max value : 433.94
# plot raster
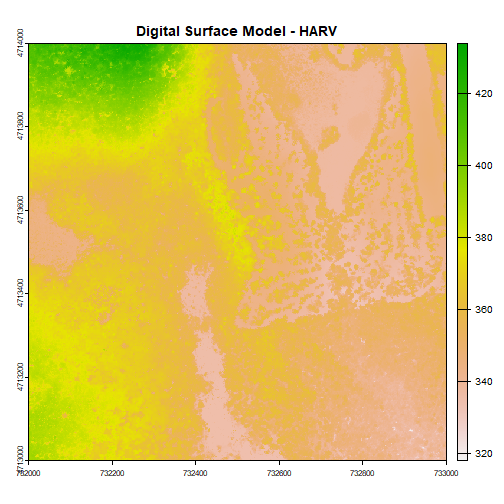
plot(DSM_HARV, main="Digital Surface Model - HARV")
Types of Data Stored in Raster Format
Raster data can be continuous or categorical. Continuous rasters can have a range of quantitative values. Some examples of continuous rasters include:
- Precipitation maps.
- Maps of tree height derived from LiDAR data.
- Elevation values for a region.
The raster we loaded and plotted earlier was a digital surface model, or a map of the elevation for Harvard Forest derived from the NEON AOP LiDAR sensor. Elevation is represented as a continuous numeric variable in this map. The legend shows the continuous range of values in the data from around 300 to 420 meters.
Some rasters contain categorical data where each pixel represents a discrete class such as a landcover type (e.g., "forest" or "grassland") rather than a continuous value such as elevation or temperature. Some examples of classified maps include:
- Landcover/land-use maps.
- Tree height maps classified as short, medium, tall trees.
- Elevation maps classified as low, medium and high elevation.
Categorical Landcover Map for the United States
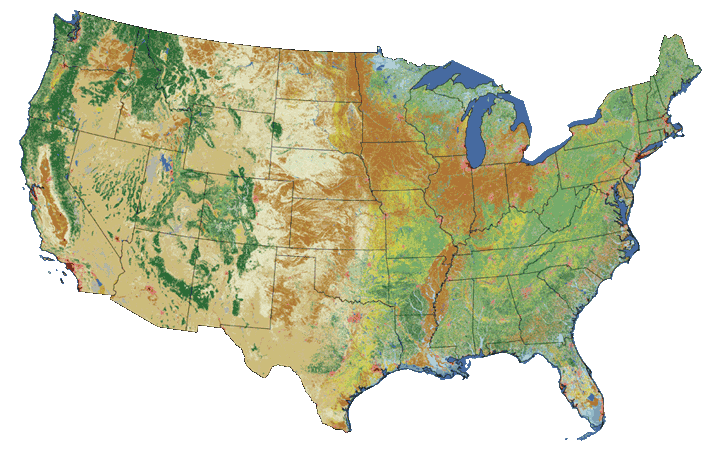
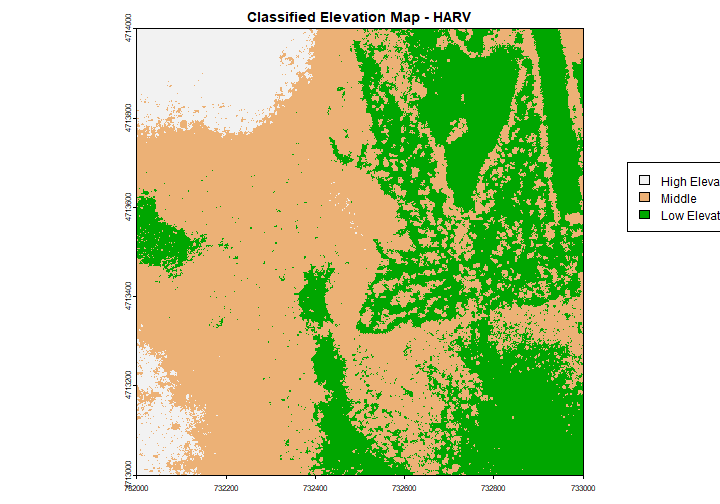
Categorical Elevation Map of the NEON Harvard Forest Site
The legend of this map shows the colors representing each discrete class.
# add a color map with 5 colors
col=terrain.colors(3)
# add breaks to the colormap (4 breaks = 3 segments)
brk <- c(250,350, 380,500)
# Expand right side of clipping rect to make room for the legend
par(xpd = FALSE,mar=c(5.1, 4.1, 4.1, 4.5))
# DEM with a custom legend
plot(DSM_HARV,
col=col,
breaks=brk,
main="Classified Elevation Map - HARV",
legend = FALSE
)
# turn xpd back on to force the legend to fit next to the plot.
par(xpd = TRUE)
# add a legend - but make it appear outside of the plot
legend( 733100, 4713700,
legend = c("High Elevation", "Middle","Low Elevation"),
fill = rev(col))
What is a GeoTIFF??
Raster data can come in many different formats. In this tutorial, we will use the
geotiff format which has the extension .tif. A .tif file stores metadata
or attributes about the file as embedded tif tags. For instance, your camera
might store a tag that describes the make and model of the camera or the date the
photo was taken when it saves a .tif. A GeoTIFF is a standard .tif image
format with additional spatial (georeferencing) information embedded in the file
as tags. These tags can include the following raster metadata:
- A Coordinate Reference System (
CRS) - Spatial Extent (
extent) - Values that represent missing data (
NoDataValue) - The
resolutionof the data
In this tutorial we will discuss all of these metadata tags.
More about the .tif format:
Coordinate Reference System
The Coordinate Reference System or CRS tells R where the raster is located
in geographic space. It also tells R what method should be used to "flatten"
or project the raster in geographic space.
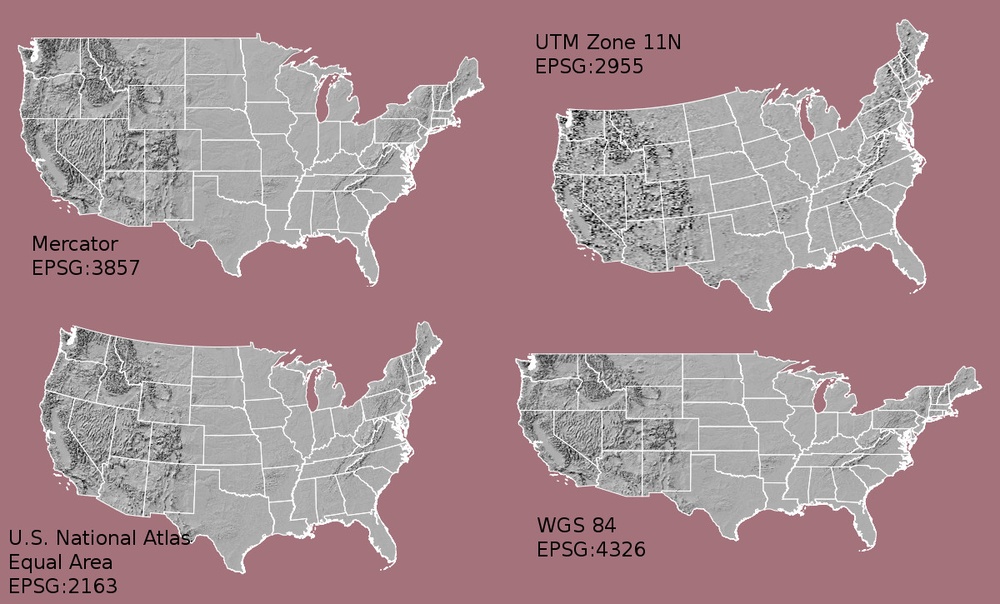
What Makes Spatial Data Line Up On A Map?
There are many great resources that describe coordinate reference systems and projections in greater detail (read more, below). For the purposes of this activity, it is important to understand that data from the same location but saved in different projections will not line up in any GIS or other program. Thus, it's important when working with spatial data in a program like R to identify the coordinate reference system applied to the data and retain it throughout data processing and analysis.
Read More:
- A comprehensive online library of CRS information.
- QGIS Documentation - CRS Overview.
- Choosing the Right Map Projection.
- NCEAS Overview of CRS in R.
How Map Projections Can Fool the Eye
Check out this short video, from Buzzfeed, highlighting how map projections can make continents seems proportionally larger or smaller than they actually are!
View Raster Coordinate Reference System (CRS) in R
We can view the CRS string associated with our R object using thecrs()
method. We can assign this string to an R object, too.
# view crs description
crs(DSM_HARV,describe=TRUE)
## name authority code
## 1 WGS 84 / UTM zone 18N EPSG 32618
## area
## 1 Between 78°W and 72°W, northern hemisphere between equator and 84°N, onshore and offshore. Bahamas. Canada - Nunavut; Ontario; Quebec. Colombia. Cuba. Ecuador. Greenland. Haiti. Jamaica. Panama. Turks and Caicos Islands. United States (USA). Venezuela
## extent
## 1 -78, -72, 84, 0
# assign crs to an object (class) to use for reprojection and other tasks
harvCRS <- crs(DSM_HARV)
The CRS of our DSM_HARV object tells us that our data are in the UTM projection, in zone 18N.

The CRS in this case is in a char format. This means that the projection
information is strung together as a series of text elements.
We'll focus on the first few components of the CRS, as described above.
-
name: The projection of the dataset. Our data are in WGS84 (World Geodetic System 1984) / UTM (Universal Transverse Mercator) zone 18N. WGS84 is the datum. The UTM projection divides up the world into zones, this element tells you which zone the data are in. Harvard Forest is in Zone 18. -
authority: EPSG (European Petroleum Survey Group) - organization that maintains a geodetic parameter database with standard codes -
code: The EPSG code. For more details, see EPSG 32618.
Extent
The spatial extent is the geographic area that the raster data covers.
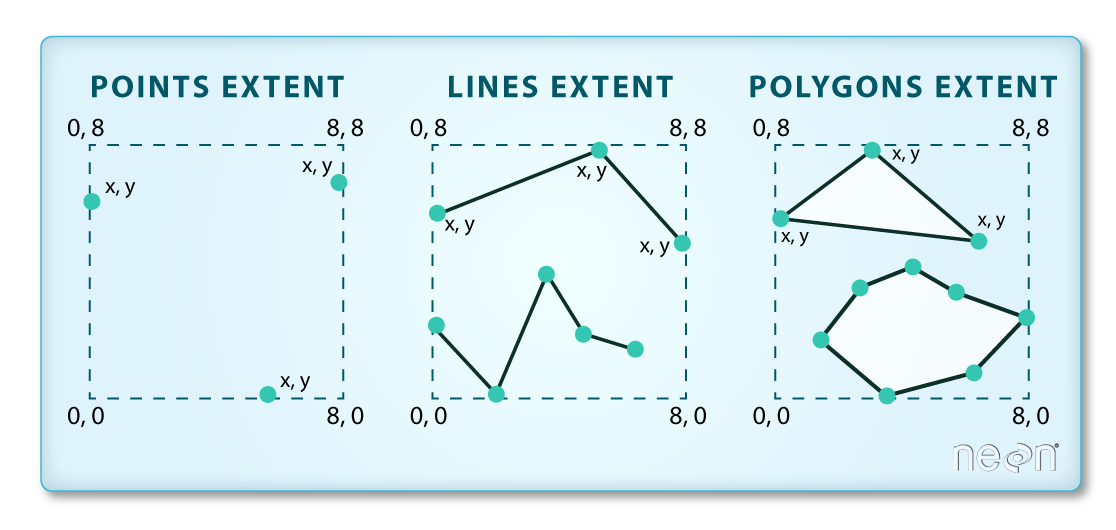
The spatial extent of an R spatial object represents the geographic "edge" or
location that is the furthest north, south, east and west. In other words, extent
represents the overall geographic coverage of the spatial object.
You can see the spatial extent using terra::ext:
ext(DSM_HARV)
## SpatExtent : 732000, 733000, 4713000, 4714000 (xmin, xmax, ymin, ymax)
Resolution
A raster has horizontal (x and y) resolution. This resolution represents the area on the ground that each pixel covers. The units for our data are in meters. Given our data resolution is 1 x 1, this means that each pixel represents a 1 x 1 meter area on the ground.

The best way to view resolution units is to look at the coordinate reference system string crs(rast,proj=TRUE). Notice our data contains: +units=m.
crs(DSM_HARV,proj=TRUE)
## [1] "+proj=utm +zone=18 +datum=WGS84 +units=m +no_defs"
Display Raster Min and Max Values
It can be useful to know the minimum or maximum values of a raster dataset. In this case, given we are working with elevation data, these values represent the min/max elevation range at our site.
Raster statistics are often calculated and embedded in a Geotiff for us.
However if they weren't already calculated, we can calculate them using the
min() or max() functions.
# view the min and max values
min(DSM_HARV)
## class : SpatRaster
## dimensions : 1000, 1000, 1 (nrow, ncol, nlyr)
## resolution : 1, 1 (x, y)
## extent : 732000, 733000, 4713000, 4714000 (xmin, xmax, ymin, ymax)
## coord. ref. : WGS 84 / UTM zone 18N (EPSG:32618)
## source(s) : memory
## varname : NEON_D01_HARV_DP3_732000_4713000_DSM
## name : min
## min value : 317.91
## max value : 433.94
We can see that the elevation at our site ranges from 317.91 m to 433.94 m.
NoData Values in Rasters
Raster data often has a NoDataValue associated with it. This is a value
assigned to pixels where data are missing or no data were collected.
By default the shape of a raster is always square or rectangular. So if we
have a dataset that has a shape that isn't square or rectangular, some pixels
at the edge of the raster will have NoDataValues. This often happens when the
data were collected by an airplane which only flew over some part of a defined
region.
Let's take a look at some of the RGB Camera data over HARV, this time downloading a tile at the edge of the flight box.
byTileAOP(dpID='DP3.30010.001',
site='HARV',
year='2022',
easting=737500,
northing=4701500,
check.size=FALSE, # set to TRUE or remove if you want to check the size before downloading
savepath = wd)
This file will be downloaded into a nested subdirectory under the ~/data folder, inside a folder named DP3.30010.001 (the Camera Data Product ID). The file should show up in this location: ~/data/DP3.30010.001/neon-aop-products/2022/FullSite/D01/2022_HARV_7/L3/Camera/Mosaic/2022_HARV_7_737000_4701000_image.tif.
In the image below, the pixels that are black have NoDataValues. The camera did not collect data in these areas.
# Use rast function to read in all bands
RGB_HARV <-
rast(paste0(wd,"DP3.30010.001/neon-aop-products/2022/FullSite/D01/2022_HARV_7/L3/Camera/Mosaic/2022_HARV_7_737000_4701000_image.tif"))
# Create an RGB image from the raster
par(col.axis="white",col.lab="white",tck=0)
plotRGB(RGB_HARV, r = 1, g = 2, b = 3, axes=TRUE)
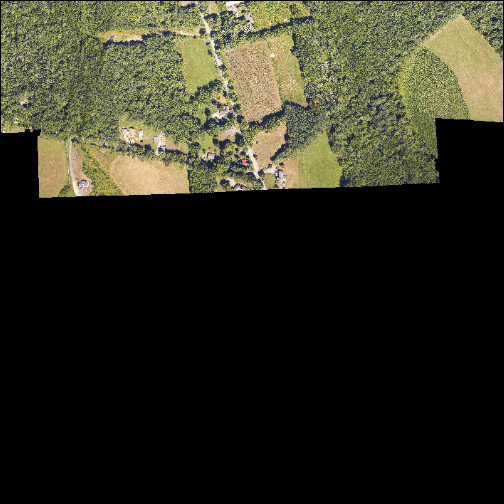

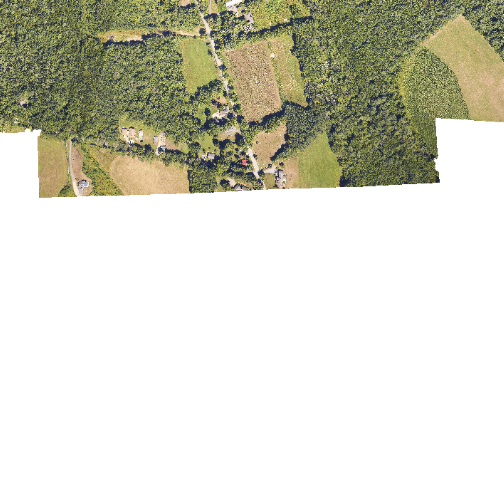
In the next image, the black edges have been assigned NoDataValue. R doesn't
render pixels that contain a specified NoDataValue. R assigns missing data
with the NoDataValue as NA.
# reassign cells with 0,0,0 to NA
func <- function(x) {
x[rowSums(x == 0) == 3, ] <- NA
x}
newRGBImage <- app(RGB_HARV, func)
##
|---------|---------|---------|---------|
par(col.axis="white",col.lab="white",tck=0)
# Create an RGB image from the raster stack
plotRGB(newRGBImage, r = 1, g = 2, b = 3, axis=TRUE)
## Warning in plot.window(...): "axis" is not a graphical parameter
## Warning in plot.xy(xy, type, ...): "axis" is not a graphical parameter
## Warning in title(...): "axis" is not a graphical parameter

NoData Value Standard
The assigned NoDataValue varies across disciplines; -9999 is a common value
used in both the remote sensing field and the atmospheric fields. It is also
the standard used by the
National Ecological Observatory Network (NEON).
If we are lucky, our GeoTIFF file has a tag that tells us what is the
NoDataValue. If we are less lucky, we can find that information in the
raster's metadata. If a NoDataValue was stored in the GeoTIFF tag, when R
opens up the raster, it will assign each instance of the value to NA. Values
of NA will be ignored by R as demonstrated above.
Bad Data Values in Rasters
Bad data values are different from NoDataValues. Bad data values are values
that fall outside of the applicable range of a dataset.
Examples of Bad Data Values:
- The normalized difference vegetation index (NDVI), which is a measure of greenness, has a valid range of -1 to 1. Any value outside of that range would be considered a "bad" value.
- Reflectance data in an image should range from 0-1 (or 0-10,000 depending upon how the data are scaled). Thus a value greater than 1 or greater than 10,000 is likely caused by an error in either data collection or processing. These erroneous values can occur, for example, in water vapor absorption bands, which contain invalid data, and are meant to be disregarded.
Find Bad Data Values
Sometimes a raster's metadata will tell us the range of expected values for a raster. Values outside of this range are suspect and we need to consider than when we analyze the data. Sometimes, we need to use some common sense and scientific insight as we examine the data - just as we would for field data to identify questionable values.
Create A Histogram of Raster Values
We can explore the distribution of values contained within our raster using the
hist() function which produces a histogram. Histograms are often useful in
identifying outliers and bad data values in our raster data.
# view histogram of data
hist(DSM_HARV,
main="Distribution of Digital Surface Model Values\n NEON Harvard Forest (HARV)",
xlab="DSM Elevation Value (m)",
ylab="Frequency",
col="lightblue")
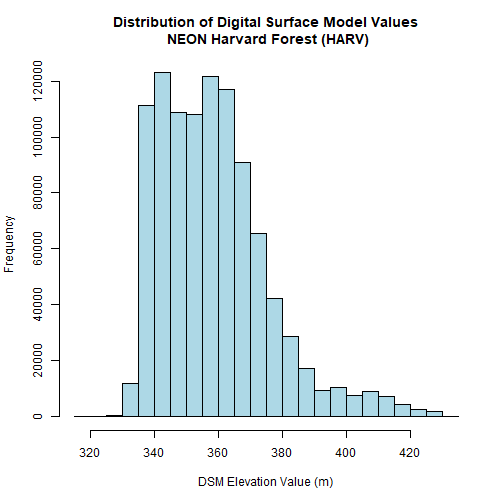
The distribution of elevation values for our Digital Surface Model (DSM) looks
reasonable. It is likely there are no bad data values in this particular raster.
Raster Bands
The Digital Surface Model object (DSM_HARV) that we've been working with
is a single band raster. This means that there is only one dataset stored in
the raster: surface elevation in meters for one time period.
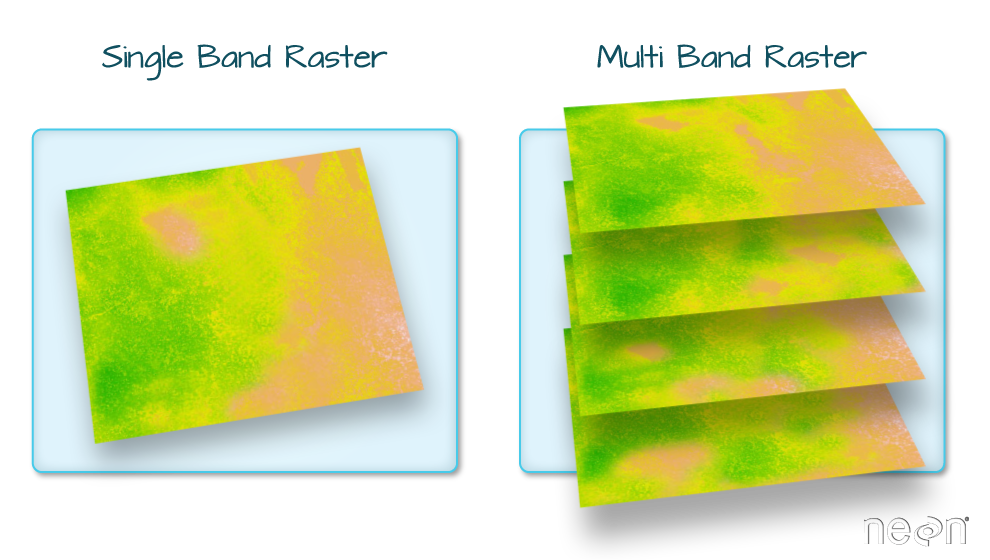

A raster dataset can contain one or more bands. We can use the rast()function to import all bands from a single OR multi-band raster. We can view the number of bands in a raster using thenlyr()` function.
# view number of bands in the Lidar DSM raster
nlyr(DSM_HARV)
## [1] 1
# view number of bands in the RGB Camera raster
nlyr(RGB_HARV)
## [1] 3
As we see from the RGB camera raster, raster data can also be multi-band,
meaning one raster file contains data for more than one variable or time period
for each cell. By default the terra::rast() function imports all bands of a
multi-band raster. You can set lyrs = 1 if you only want to read in the first
layer, for example.
Jump to the fourth tutorial in this series for a tutorial on multi-band rasters: Work with Multi-band Rasters: Images in R.
View Raster File Attributes
Remember that a GeoTIFF contains a set of embedded tags that contain
metadata about the raster. So far, we've explored raster metadata after
importing it in R. However, we can use the describe("path-to-raster-here")
function to view raster information (such as metadata) before we open a file in
R. Use help(describe) to see other options for exploring the file contents.
# view metadata attributes before opening the file
describe(path.expand(dsm_harv_file),meta=TRUE)
## [1] "AREA_OR_POINT=Area"
## [2] "TIFFTAG_ARTIST=Created by the National Ecological Observatory Network (NEON)"
## [3] "TIFFTAG_COPYRIGHT=The National Ecological Observatory Network is a project sponsored by the National Science Foundation and managed under cooperative agreement by Battelle. This material is based in part upon work supported by the National Science Foundation under Grant No. DBI-0752017."
## [4] "TIFFTAG_DATETIME=Flown on 2022080312, 2022080412, 2022081213, 2022081413"
## [5] "TIFFTAG_IMAGEDESCRIPTION=Elevation LiDAR - NEON.DP3.30024 acquired at HARV by RIEGL LASER MEASUREMENT SYSTEMS Q780 2220855 as part of 2022-P3C1"
## [6] "TIFFTAG_MAXSAMPLEVALUE=434"
## [7] "TIFFTAG_MINSAMPLEVALUE=318"
## [8] "TIFFTAG_RESOLUTIONUNIT=2 (pixels/inch)"
## [9] "TIFFTAG_SOFTWARE=Tif file created with a Matlab script (write_gtiff.m) written by Tristan Goulden (tgoulden@battelleecology.org) with data processed from the following scripts: create_tiles_from_mosaic.m, combine_dtm_dsm_gtif.m, lastools_workflow.csh which implemented LAStools version 210418."
## [10] "TIFFTAG_XRESOLUTION=1"
## [11] "TIFFTAG_YRESOLUTION=1"
Specifying options=c("stats") will show some summary statistics:
# view summary statistics before opening the file
describe(path.expand(dsm_harv_file),options=c("stats"))
## [1] "Driver: GTiff/GeoTIFF"
## [2] "Files: C:/Users/bhass/Documents/data/DP3.30024.001/neon-aop-products/2022/FullSite/D01/2022_HARV_7/L3/DiscreteLidar/DSMGtif/NEON_D01_HARV_DP3_732000_4713000_DSM.tif"
## [3] " C:/Users/bhass/Documents/data/DP3.30024.001/neon-aop-products/2022/FullSite/D01/2022_HARV_7/L3/DiscreteLidar/DSMGtif/NEON_D01_HARV_DP3_732000_4713000_DSM.tif.aux.xml"
## [4] "Size is 1000, 1000"
## [5] "Coordinate System is:"
## [6] "PROJCRS[\"WGS 84 / UTM zone 18N\","
## [7] " BASEGEOGCRS[\"WGS 84\","
## [8] " ENSEMBLE[\"World Geodetic System 1984 ensemble\","
## [9] " MEMBER[\"World Geodetic System 1984 (Transit)\"],"
## [10] " MEMBER[\"World Geodetic System 1984 (G730)\"],"
## [11] " MEMBER[\"World Geodetic System 1984 (G873)\"],"
## [12] " MEMBER[\"World Geodetic System 1984 (G1150)\"],"
## [13] " MEMBER[\"World Geodetic System 1984 (G1674)\"],"
## [14] " MEMBER[\"World Geodetic System 1984 (G1762)\"],"
## [15] " MEMBER[\"World Geodetic System 1984 (G2139)\"],"
## [16] " ELLIPSOID[\"WGS 84\",6378137,298.257223563,"
## [17] " LENGTHUNIT[\"metre\",1]],"
## [18] " ENSEMBLEACCURACY[2.0]],"
## [19] " PRIMEM[\"Greenwich\",0,"
## [20] " ANGLEUNIT[\"degree\",0.0174532925199433]],"
## [21] " ID[\"EPSG\",4326]],"
## [22] " CONVERSION[\"UTM zone 18N\","
## [23] " METHOD[\"Transverse Mercator\","
## [24] " ID[\"EPSG\",9807]],"
## [25] " PARAMETER[\"Latitude of natural origin\",0,"
## [26] " ANGLEUNIT[\"degree\",0.0174532925199433],"
## [27] " ID[\"EPSG\",8801]],"
## [28] " PARAMETER[\"Longitude of natural origin\",-75,"
## [29] " ANGLEUNIT[\"degree\",0.0174532925199433],"
## [30] " ID[\"EPSG\",8802]],"
## [31] " PARAMETER[\"Scale factor at natural origin\",0.9996,"
## [32] " SCALEUNIT[\"unity\",1],"
## [33] " ID[\"EPSG\",8805]],"
## [34] " PARAMETER[\"False easting\",500000,"
## [35] " LENGTHUNIT[\"metre\",1],"
## [36] " ID[\"EPSG\",8806]],"
## [37] " PARAMETER[\"False northing\",0,"
## [38] " LENGTHUNIT[\"metre\",1],"
## [39] " ID[\"EPSG\",8807]]],"
## [40] " CS[Cartesian,2],"
## [41] " AXIS[\"(E)\",east,"
## [42] " ORDER[1],"
## [43] " LENGTHUNIT[\"metre\",1]],"
## [44] " AXIS[\"(N)\",north,"
## [45] " ORDER[2],"
## [46] " LENGTHUNIT[\"metre\",1]],"
## [47] " USAGE["
## [48] " SCOPE[\"Navigation and medium accuracy spatial referencing.\"],"
## [49] " AREA[\"Between 78°W and 72°W, northern hemisphere between equator and 84°N, onshore and offshore. Bahamas. Canada - Nunavut; Ontario; Quebec. Colombia. Cuba. Ecuador. Greenland. Haiti. Jamaica. Panama. Turks and Caicos Islands. United States (USA). Venezuela.\"],"
## [50] " BBOX[0,-78,84,-72]],"
## [51] " ID[\"EPSG\",32618]]"
## [52] "Data axis to CRS axis mapping: 1,2"
## [53] "Origin = (732000.000000000000000,4714000.000000000000000)"
## [54] "Pixel Size = (1.000000000000000,-1.000000000000000)"
## [55] "Metadata:"
## [56] " AREA_OR_POINT=Area"
## [57] " TIFFTAG_ARTIST=Created by the National Ecological Observatory Network (NEON)"
## [58] " TIFFTAG_COPYRIGHT=The National Ecological Observatory Network is a project sponsored by the National Science Foundation and managed under cooperative agreement by Battelle. This material is based in part upon work supported by the National Science Foundation under Grant No. DBI-0752017."
## [59] " TIFFTAG_DATETIME=Flown on 2022080312, 2022080412, 2022081213, 2022081413"
## [60] " TIFFTAG_IMAGEDESCRIPTION=Elevation LiDAR - NEON.DP3.30024 acquired at HARV by RIEGL LASER MEASUREMENT SYSTEMS Q780 2220855 as part of 2022-P3C1"
## [61] " TIFFTAG_MAXSAMPLEVALUE=434"
## [62] " TIFFTAG_MINSAMPLEVALUE=318"
## [63] " TIFFTAG_RESOLUTIONUNIT=2 (pixels/inch)"
## [64] " TIFFTAG_SOFTWARE=Tif file created with a Matlab script (write_gtiff.m) written by Tristan Goulden (tgoulden@battelleecology.org) with data processed from the following scripts: create_tiles_from_mosaic.m, combine_dtm_dsm_gtif.m, lastools_workflow.csh which implemented LAStools version 210418."
## [65] " TIFFTAG_XRESOLUTION=1"
## [66] " TIFFTAG_YRESOLUTION=1"
## [67] "Image Structure Metadata:"
## [68] " INTERLEAVE=BAND"
## [69] "Corner Coordinates:"
## [70] "Upper Left ( 732000.000, 4714000.000) ( 72d10'28.52\"W, 42d32'36.84\"N)"
## [71] "Lower Left ( 732000.000, 4713000.000) ( 72d10'29.98\"W, 42d32' 4.46\"N)"
## [72] "Upper Right ( 733000.000, 4714000.000) ( 72d 9'44.73\"W, 42d32'35.75\"N)"
## [73] "Lower Right ( 733000.000, 4713000.000) ( 72d 9'46.20\"W, 42d32' 3.37\"N)"
## [74] "Center ( 732500.000, 4713500.000) ( 72d10' 7.36\"W, 42d32'20.11\"N)"
## [75] "Band 1 Block=1000x1 Type=Float32, ColorInterp=Gray"
## [76] " Min=317.910 Max=433.940 "
## [77] " Minimum=317.910, Maximum=433.940, Mean=358.584, StdDev=17.156"
## [78] " NoData Value=-9999"
## [79] " Metadata:"
## [80] " STATISTICS_MAXIMUM=433.94000244141"
## [81] " STATISTICS_MEAN=358.58371301653"
## [82] " STATISTICS_MINIMUM=317.91000366211"
## [83] " STATISTICS_STDDEV=17.156044149253"
## [84] " STATISTICS_VALID_PERCENT=100"
It can be useful to use describe to explore your file before reading it into R.
Challenge: Explore Raster Metadata
Without using the terra function to read the file into R, determine the following information about the DTM file. This was downloaded at the same time as the DSM file, and as long as you didn't move the data, it should be located here: ~/data/DP3.30024.001/neon-aop-products/2022/FullSite/D01/2022_HARV_7/L3/DiscreteLidar/DTMGtif/NEON_D01_HARV_DP3_732000_4713000_DTM.tif.
- Does this file has the same
CRSasDSM_HARV? - What is the
NoDataValue? - What is resolution of the raster data?
- Is the file a multi- or single-band raster?
- On what dates was this site flown?